Karen Eilbeck
Professor of Biomedical Informatics and Adjunct Associate Professor of Human Genetics
Bioinformatics Research

Molecular Biology Program
Education
B.S. University of Salford, United Kingdom
Ph.D. University of Manchester, United Kingdom
Research
The lab performs bioinformatics research. We are interested in using aspects of computer science to address contemporary genomics and molecular biology questions, with a focus on precision medicine. We are interested in the management and analysis of biological data using bioinformatics techniques. The biological sciences are data intensive but how the data is organized affects how well it can be analyzed and shared with the community. Ontologies are used to standardize and structure the data to make it more amenable to analysis and allow the user community to communicate with each other using the same language. An ontology is a description of our knowledge about a subject. It contains the classes of the data and the relationships that hold between them. The focus of the lab is to develop ontologies and ontology-enabled software to enhance biological research—doing so is essential for analyzing and managing the kinds of genomic and biological data that exist today.
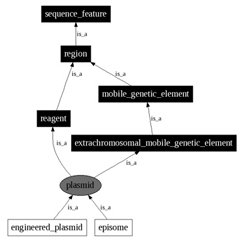
User interface to query the Sequence Ontology
References (Selected Publications)
- Henrie A, Hemphill SE, Ruiz-Schultz N, Cushman B, DiStefano MT, Azzariti D, Harrison SM, Rehm HL, Eilbeck K. (2018) ClinVar Miner: Demonstrating utility of a Web-based tool for viewing and filtering ClinVar data. Hum Mutat 39(8), 1051-1060
- Eilbeck K, Quinlan A, Yandell M. (2017) Settling the score: Variant prioritization and Mendelian disease. Nat Rev Genet18(10), 599-612
- Schlaberg R, Queen K, Simmon K, Tardif K, Stockmann C, Flygare S, Kennedy B, Voelkerding K, Bramley A, Zhang J, Eilbeck K, Yandell M, Jain S, Pavia AT, Tong S, Ampofo K. (2017) Viral Pathogen Detection by Metagenomics and Pan-Viral Group Polymerase Chain Reaction in Children With Pneumonia Lacking Identifiable Etiology.J Infect Dis215(9), 1407-1415
- Lubin IM, Aziz N, Babb LJ, Ballinger D, Bisht H, Church DM, Cordes S, Eilbeck K, Hyland F, Kalman L, Landrum M, Lockhart ER, Maglott D, Marth G, Pfeifer JD, Rehm HL, Roy S, Tezak Z, Truty R, Ullman-Cullere M, Voelkerding KV, Worthey EA, Zaranek AW, Zook JM. (2017) Principles and Recommendations for Standardizing the Use of the Next-Generation Sequencing Variant File in Clinical Settings.J Mol Diagn19(3), 417-426.
- Flygare S, Simmon K, Miller C, Qiao Y, Kennedy B, Di Sera T, Graf EH, Tardif KD, Kapusta A, Rynearson S, Stockmann C, Queen K, Tong S, Voelkerding KV, Blaschke A, Byington CL, Jain S, Pavia A, Ampofo K, Eilbeck K, Marth G, Yandell M, Schlaberg R. (2016) Taxonomer: an interactive metagenomics analysis portal for universal pathogen detection and host mRNA expression profiling. Genome Biol 17(1), 111.
- Huang J, Gutierrez F, Strachan HJ, Dou D, Huang W, Smith B, Blake JA, Eilbeck K, Natale DA, Lin Y, Wu B, Silva Nd, Wang X, Liu Z, Borchert GM, Tan M, Ruttenberg A. (2016) OmniSearch: a semantic search system based on the Ontology for MIcroRNA Target (OMIT) for microRNA-target gene interaction data.J Biomed Semantics 7, 25.
- Huang J, Eilbeck K, Smith B, Blake JA, Dou D, Huang W, Natale DA, Ruttenberg A, Huan J, Zimmermann MT, Jiang G, Lin Y, Wu B, Strachan HJ, He Y, Zhang S, Wang X, Liu Z, Borchert GM, Tan M. (2016) The Non-Coding RNA Ontology (NCRO): a comprehensive resource for the unification of non-coding RNA biology. J Biomed Semantics7, 24.
- Graf EH, Simmon KE, Tardif KD, Hymas W, Flygare S, Eilbeck K, Yandell M, Schlaberg R. (2016) Unbiased Detection of Respiratory Viruses by Use of RNA Sequencing-Based Metagenomics: a Systematic Comparison to a Commercial PCR Panel. J Clin Microbiol54(4), 1000-7.
- Desvignes T, Batzel P, Berezikov E, Eilbeck K, Eppig JT, McAndrews MS, Singer A, Postlethwait JH. (2015) miRNA Nomenclature: A View Incorporating Genetic Origins, Biosynthetic Pathways, and Sequence Variants. Trends Genet31(11), 613-26.
- Duncan J, Eilbeck K, Narus SP, Clyde S, Thornton S, Staes C. (2015) Building an Ontology for Identity Resolution in Healthcare and Public Health. Online J Public Health Inform7(2), e219. Cunningham F, Moore B, Ruiz-Schultz N, Ritchie GR, Eilbeck K. (2015) Improving the Sequence Ontology terminology for genomic variant annotation.J Biomed Semantics 6, 32.
- Singleton MV, Guthery SL, Voelkerding KV, Chen K, Kennedy B, Margraf RL, Durtschi J, Eilbeck K, Reese MG, Jorde LB, Huff CD, Yandell M. (2014) Phevor combines multiple biomedical ontologies for accurate identification of disease-causing alleles in single individuals and small nuclear families. Am J Hum Genet94(4), 599-610.
- Reese MG, Moore B, Batchelor C, Salas F, Cunningham F, Marth GT, Stein L, Flicek P, Yandell M, Eilbeck K. (2010). A standard variation file format for human genome sequences. Genome Biol 11(8), R88.
- Eilbeck K, Moore B, Holt C and Yandell M. (2009) Quantitative measures for the management and comparison of annotated genomes. BMC Bioinformatics10, 67.
- Eilbeck K, Lewis SE, Mungall CJ, Yandell M, Stein L, Durbin R, Ashburner M. (2005) The Sequence Ontology: a tool for the unification of genome annotations.Genome Biology. 6(5), R44.
- Venter JC, et al. (2001) The sequence of the human genome.Science291(5507), 1304-51. Erratum in: Science 2001 Jun 5; 292(5523), 1838.
